Phred is a base calling program for DNA sequence traces; Phred executables for Windows, Mac OS X, Linux, and Unix are available from CodonCode Corporation as part of the PHRED - PHRAP package.
- Sanger Sequence Assembly software, free download Mac Os X
- Sanger Sequence Assembly software, free download Mac Os Bootable
Phred was developed by Drs. Phil Green and Brent Ewing, and is distributed by CodonCode Corporation under license from the University of Washington. Phred is widely used by the largest academic and commercial DNA sequencing laboratories. This page gives a brief description of Phred. For information about Phrap, Cross_match, and Consed, please visit www.phrap.com.
In bioinformatics, sequence assembly refers to aligning and merging fragments from a longer DNA sequence in order to reconstruct the original sequence. This is needed as DNA sequencing technology cannot read whole genomes in one go, but rather reads small pieces of between 20 and 30,000 bases, depending on the technology used. Typically the short fragments, called reads, result from shotgun. BioEdit - a very popular free sequence alignment editor for Windows. Staden Package - a powerful open source sequence assembly and editing package for UNIX, Linux, Windows, and Mac OS X. Not the most user-friendly package, steep learning curve. Se-Al - An older sequence alignment editor for Mac OS X. Not updated since 2002, but.
Since Phred was developed for easy integration into automated data processing pipelines, Phred does not provide a graphical user interface. For scientists who would like to use Phred on Windows or Mac OS X from a user-friendly graphical interface, CodonCode Corporation offers CodonCode Aligner.
Sanger Sequence Assembly software, free download Mac Os X
What Phred Does
Phred is a base-calling program for DNA sequence traces. Phred reads DNA sequence chromatogram files and analyzes the peaks to call bases, assigning quality scores ('Phred scores') to each base call.
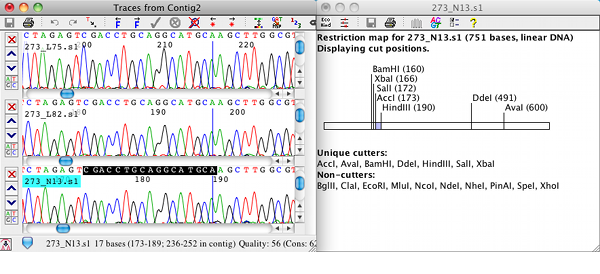

Phred can read input files in the following formats:
- SCF ('Standard Chromatogram Format') files. The SCF format is a 'universal' format for trace sequence files that is supported by many different programs and manufacturers.
- Applied Biosystems (ABI) chromatogram files.
Phred can produce a variety of different output files:
- SCF files with Phred's base calls and quality values
- Sequence files in FASTA (or XBAP) format
- Quality files in FASTA (or XPAB) format
- PHD files - text files which contain base call and quality information, which can later be used during contig editing by Consed and similar programs.
One good reason to use Phred for base calling is higher accuracy: in one study, Phred made '40-50% fewer errors' 'than the ABI software' (Ewing et al.1998a, Genome Research 8:175-85 ). Since this initial study, ABI has improved their base calling software, and eventually incorporated base-specific quality scores similar to Phred scores into their 'KB' base caller.
Another very interesting feature of Phred is the generation of highly accurate, base-specific quality scores (see next section). Phred quality scores have become widely accepted to characterize the quality of sequences, for example to compare different sequencing methods. They are also used by Phil Green's sequence assembly program Phrap to generate better assemblies; how Phred works together with Phrap is described below.
Phred Quality Scores
Phred's base-specific quality scores are one of the most innovative features in Phred. After calling bases, Phred examines the peaks around each base call to assign a quality score to each base call. Quality scores range from 4 to about 60, with higher values corresponding to higher quality. The quality scores are logarithmically linked to error probabilities, as shown in the following table:
Phred quality score | Probability that the base is called wrong | Accuracy of the base call |
|---|---|---|
10 | 1 in 10 | 90% |
20 | 1 in 100 | 99% |
30 | 1 in 1,000 | 99.9% |
40 | 1 in 10,000 | 99.99% |
50 | 1 in 100,000 | 99.999% |
It has been shown that Phred's error probabilities are very accurate - if Phred assigns a quality score of 40 to a base, the chances that this base is called incorrectly are indeed just 1 in 10,000 (see Ewing et al.1998b, Genome Research 8:186-94 ). This high accuracy has been observed for sequences generated at different laboratories, each using a different combination of sequencing enzymes, fluorescent dyes, and gel run conditions (Richterich 1998, Genome Research; 8:251-9).
The high accuracy of Phred quality scores make them an ideal tool to assess the quality of sequences. The most commonly used method is to count the bases with a quality score of 20 and above (sometimes called 'high quality bases'); the resulting number is often called the 'Phred20 score'. By looking at individual sequences, failed reactions or low-quality reads can easily be identified. When looking at collections of sequences, the effect of different sequencing methods on sequence quality can be directly measured. This allows straighforward quality control in sequencing projects, and can give easily available measures to optimize sequencing operations.
Support for Phred quality scores is fully integrated into CodonCode Aligner. Aligner enables you to run Phred on sequence traces by simply selecting you sequences and choosing 'Call Bases' from a menu. CodonCode Aligner can show Phred quality scores in a number of different ways: by shading the background behind bases according to quality, in a separate 'quality view', or as a summary of 'Phred20' scores in the project view. CodonCode Aligner can also use quality scores during sequence assembly to build the consensus sequence, automatically selecting the consensus base that is most likely to be correct. Such quality-based consensus sequences can be much more accurate than majority-based sequences, especially in areas of low coverage.
Free demo versions of CodonCode Aligner are available for download. The demo version of CodonCode Aligner includes special 'workstation' versions of Phred and Phrap, which can be used after requesting a trial license or purchasing a license for CodonCode Aligner (the workstation versions are identical to the regular programs, except that they can be run only from CodonCode Aligner).
Phred, Phrap, and Cross_match
Phred is part of a larger set of programs for DNA sequencing, all of which were developed in Dr. Phil Green's group. In most sequencing projects, Phred is used in together with Cross_match for vector screening, and Phrap for sequence assembly. Phrap uses Phred's quality values in several ways:
- Better identification of repeat sequences: If two overlapping sequences have high quality discrepancies, Phrap can deduce that such difference are caused by sequencing different copies of repeats (rather than random errors), and place the reads in two different contigs.
- Better 'consensus' sequences. Phrap builds consensus sequences by picking the highest quality read at each position in a contig, very much like human 'contig editors' do. Phrap also looks at all individual reads at each position to assign a quality value to the consensus, taking discrepancies and confirmations by different strands or sequencing chemistries into account.
For more information about Phrap and Cross_match, please visit www.phrap.com.
Sanger Sequence Assembly software, free download Mac Os Bootable
